Welcome to Huang's page
Title:Classification of liver tumors with CEUS based on 3D-CNN*
Introduction:
The liver cancer has the third highest mortality rate in the world. The effective treatment depends on the accurate identification of benign and malignant tumors. This paper proposed a computer aided system for distinguishing liver lesions based on CEUS, a widely accepted inspection technique. Video sequences are handled by the 3D convolutional neural network (3D-CNN) to extract spatial and temporal features. Meanwhile, the framework is trained by a specific dataset to yield a classification network. According to the results, our system obtained higher performance than previous system, in which the average accuracy rate reached 93.1% with ten-fold cross-validation. It is noteworthy that the system is potential and easy to expand to other applications.
In 2D images, the local patterns such as texture and edge were integrated by 2D convolution kernel from previous feature maps. In order to extend such function into 3D space, the sliding window of convolution should be appended to temporal dimension, in which the value at position (x,y,z) was calculated as follows:
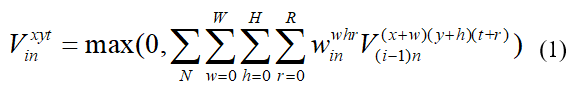
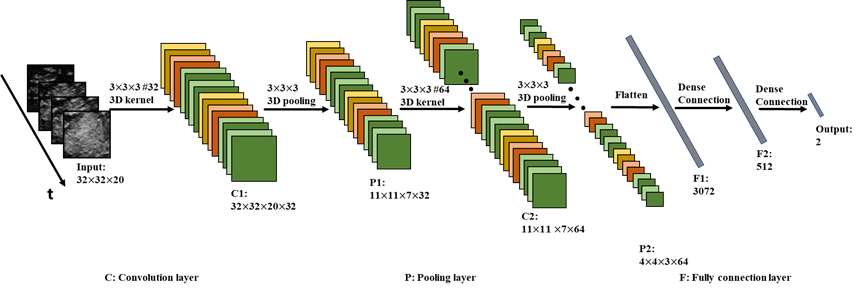
Where max(·)was Rectified Linear Unit (ReLU) activation function.i,n represented the index of convolution layer and feature map, respectively. N was the number of feature maps, and (W,H,R) was the size of voxel at position (x,y,t). Fig. 1 explained how 3D convolution worked on CEUS frames. The sliding direction of 2D image was the same as 2D-CNN but the sliding window was additionally applied over consecutive frames in 3D-CNN. For brevity, the convolution layer in Fig. 1 consisted of two continuous convolutions. At this stage, in order to increase feature maps, there were 32 and 64 of different kernels at C1 and C2, respectively. Shown in Fig.1, the size of feature map after convolution at C1 was 32x32x20, which was coincident with that of input data.